Biophysicists in Germany and the US have used nuclear magnetic resonance to gain an important new insight into how proteins interact.
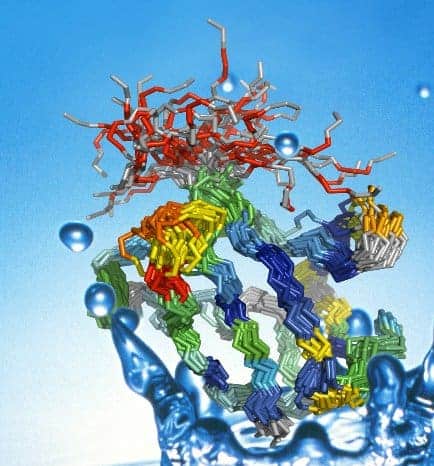
The researchers were, for the first time, able to map out all the different shapes that the protein ubiquitin can take over a period of several microseconds. The team found that many of these shapes are nearly identical to those adopted by ubiquitin when it binds with other proteins — a discovery that suggests the prevailing theory of protein binding is incomplete, and could pave the way for new types of drugs (Science 320 1471).
All living things contain proteins and many biological processes involve these chain-like molecules binding with one another. Biophysicists know that a bound protein appears to have a different shape than its free counterpart — but understanding exactly how this change in shape occurs has proven very difficult.
For more than 50 years, most biophysicists subscribed to the “induced fit” theory, whereby a free protein is coaxed by its partner to undergo a gradual change in its shape during the binding process.
Many different shapes
However, biophysicists are beginning to realize that free proteins fluctuate between many different shapes in the absence of a binding partner — and it could be that binding simply occurs when a free protein spontaneously assumes the correct shape.
This theory of “conformational selection” had been hard to establish because standard techniques such as X-ray diffraction cannot identify the large number of short-lived shapes that a free protein can adopt.
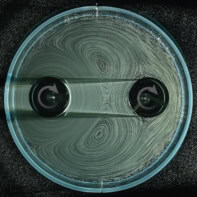
Now, Bert de Groot and colleagues at the Max Planck Institute for Biophysical Chemistry in Goettingen and Vanderbilt University in Nashville have used nuclear magnetic resonance (NMR) to map out all the possible shapes of free ubiquitin, which is a protein found in many living cells.
The team used a relatively new NMR technique, which can follow the positions of atoms in a molecule on timescales up to several microseconds. This is much longer than traditional NMR, which can only detect motion on nanosecond timescales — not long enough to get a good look at all of the possible shapes the protein can adopt, according to De Groot.
Matching 46 bound structures
The team then compared this “structural ensemble” of free protein shapes with the 46 shapes that ubiquitin is known to adopt when bound within larger structures. They discovered that every one of the bound shapes also occurred in the free protein.
De Groot told physicsworld.com that the result implies that “no induced-fit motions are required for ubiquitin to adapt to its different binding partners”.
Writing in the same issue of Science, David Boehr and Peter Wright of the Scripps Institute in California point out that the NMR work does not necessarily mean that the induced fit theory is incorrect (Science 320 1429). Rather it is possible that both mechanisms are involved in the binding process. They also point out that de Groot’s results suggest that structural fluctuations could play an important role in how the function of a protein evolves over time.
De Groot added that the findings will shed light on a number of processes that involve protein binding including biochemical signalling processes, and receptor-ligand recognition. Both of these processes are important to those designing new drugs, and de Groot believes that the team’s work could “contribute to the design of novel drugs”.