Two independent groups have used crystallography to gain insight into how some organisms use antifreeze proteins to protect against frost damage. The work suggests that water-repelling barbs on the protein’s ice-binding surface nestle into the holes at the centres of hexagonal ice crystals.
Freezing kills most organisms by rupturing their cells and vessels and therefore life in cold climates has evolved to take freezing risks seriously. Several species have independently developed antifreeze proteins that prevent this terrible fate by binding to tiny ice crystals and stopping their growth. However, researchers have struggled to understand how these proteins recognize ice.
Now a team led by Alberto Podjarny of the Institut de Génétique et de Biologie Moléculaire et Cellulaire in Strasbourg, France and Eduardo Howard of the Instituto de Física de Líquidos y Sistemas Biológicos in La Plata, Argentina has used a combination of X-ray and neutron diffraction to gain insight into the problem. Also, a group headed by Peter Davies at Queens University in Kingston, Canada, has used X-rays to learn more about natural antifreeze.
Elusive hydrogen
X-ray diffraction has long been used to help deduce the structure of proteins, giving the locations and species of the constituent atoms. However, hydrogen atoms are very difficult to spot because they only have one electron each and therefore don’t interact very strongly with X-rays. As a result it can be difficult to get an accurate protein structure using X-rays alone.
Neutrons, on the other hand, interact mostly with atomic nuclei and the single proton at the centre of a hydrogen atom scatters neutrons well. As a result hydrogen appears much more clearly in structures generated with neutron diffraction. Deuterium – hydrogen with a proton and neutron in the nucleus – is even better. “The advantage of using neutrons is that you see not only the positions of the carbon, nitrogen and oxygen atoms, but you also see the hydrogen,” says Matthew Blakeley of the Institut Laue-Langevin (ILL) in Grenoble, France who worked with Podjarny and Howard.
To make the antifreeze protein with deuterium in place of the ordinary hydrogen, the team modified E. coli bacteria to produce a form of the protein found in the ocean pout fish of the North Atlantic. The bacteria had lived in “heavy” water, with deuterium in place of the hydrogen, at ILL’s Deuteration Laboratory. This ensures that all parts of the protein would be visible to the neutron beam.
Binding problems
A large number of protein molecules are then placed in a solvent to create a crystal. When exposed to ILL’s intense neutron beam, the crystal produced the sharpest image yet of the antifreeze protein’s structure.
Crystallization is a prerequisite for such diffraction studies, but it also introduces challenges. Because of the way that the proteins bind together, the ice-binding part of the molecule is not usually exposed – making the binding process difficult to study. However, the team did find a small amount of exposed ice-binding surface, which gave them a glimpse of the protein at work.
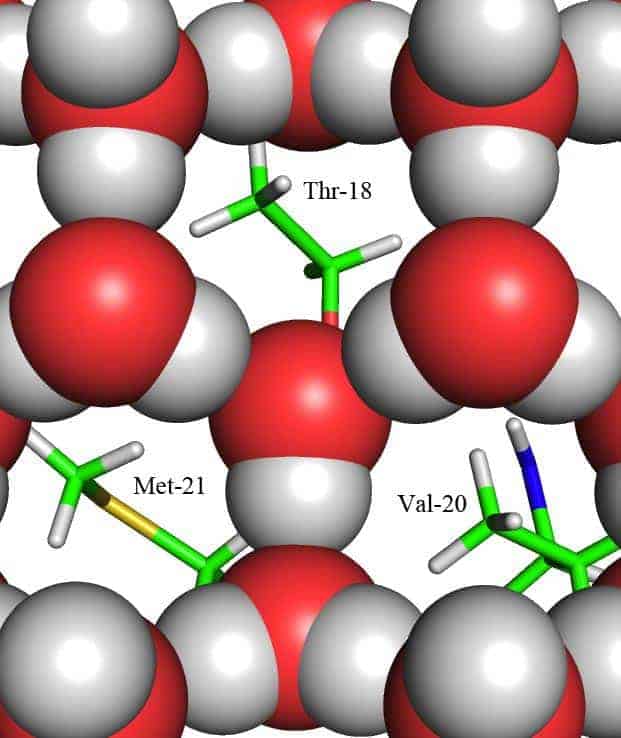
The team saw four water molecules form an ice-like tetrahedral arrangement by partly surrounding one of the hydrophobic barbs. It was easy to miss, says Blakeley, because one water molecule was mobile, occupying one of two positions. X-rays couldn’t pick it up easily, but neutrons could. Using the positions of this tetrahedral cluster, the team modelled the hexagonal ice crystal structure over the entire binding surface, with the hydrophobic barbs located at the centres of the rings.
Hexagons please
Ansgar Siemer of Columbia University in New York City, who has used nuclear magnetic resonance (NMR) to identify the ice-binding sites on antifreeze proteins, believes this detailed picture of the protein’s structure will be useful to researchers. Still, he would like to see more than four water molecules attached to the ice-binding site before saying it confirms that the spikes nestle into the hexagonal holes in ice.
That issue has been addressed by an independent team led by Peter Davies at Queen’s University. The researchers used X-rays to study of the antifreeze protein from an Antarctic bacterium. They managed to make a long swath of ice-binding surface free to attract water. On this surface, about sixty water molecules arranged themselves in an “ice-like” hexagonal formation around the water-repellent barbs.
“There is no need to model or extrapolate – we see it all very clearly,” says Davies. He suggests that these pre-attached water molecules freeze to ice crystals forming inside cells, bonding the ice to the protein.
Davies and Blakeley both suspect that the proteins distort the ice structure so that it is difficult for more water molecules to attach. Davies compares the ice to a pillow, dotted with antifreeze stones; the crystal bulges between the proteins until its curvature is too great to accept more water.
Blakeley suggests that antifreeze proteins may find application in medicine, preserving cells and tissues for later use, and controversially, they are already additives in ice cream produced by Unilever.
The ILL work is described in Journal of Molecular Recognition 24 724 and the Queen’s research in PNAS doi/10.1073/pnas.1100429108



