Two independent teams in the US have made DNA robots mimic the protein motors in our bodies – be it walking without help along predefined routes or taking cargo from A to B.
The experiments, which are the first to truly combine advances in our knowledge of DNA structure and dynamics, suggest that nanorobots could soon be performing autonomous, useful tasks.
“A goal of our field is to re-fashion and re-imagine all the complex biomechanical machinery of cells to suit our own purposes,” says Paul Rothermund, an expert in molecular robotics at the California Institute of Technology who was not involved in the research. “[We want] to have synthetic molecules that can move around, carry cargo, act as chemical factories…and above all to make these processes modular, to make them engineerable. These two papers mark a significant advance along this research direction.”
Walkers and spiders
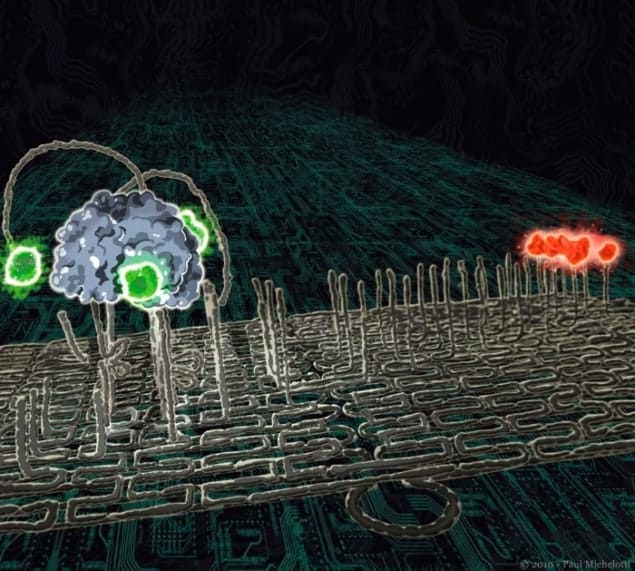
So far biophysicists have found two ways create such molecular robots. The first, known as DNA walkers, have a body and feet made from DNA, with extra “anchor” strands of DNA that join the feet to a surface. When different “fuel” strands are put in front of a walker, they preferentially join to the anchor strands, thereby freeing the walker to move forward. The second type of robot, known as a molecular spider, has a protein body and DNA legs that chemically cleave tall strands of DNA on a surface. Simply put, a molecular spider nibbles its way through a lawn of DNA, only going where there is more grass to mow.
These two papers mark a significant advance along this research direction Paul Rothermund, California Institute of Technology
But the trouble with these robots is that, in the past, they have only been shown to move slowly and aimlessly. Ideally, researchers would like them to walk in a set direction while performing a meaningful task – rather like Mother Nature’s protein motors, which move along well defined tracks in a cell while carrying cargo.
Now the two research groups – one involving Milan Stojanovic at Columbia University and colleagues, the other led by Ned Seeman at New York University – have got around this problem by exploiting developments in structural DNA, namely “DNA origami”. Based on folded DNA, this neat trick makes something akin to a pegboard to which all sorts of molecules can be attached. In the context of the current research, however, it provides an ideal track for walkers or spiders to move along.
The route to autonomous assembly
Stojanovic’s group have programmed a thin track of DNA origami with a route for a molecular spider. Using an atomic force microscope, they imaged the spider moving forwards in a straight line leaving a trail of cleaved DNA (or cut grass) behind. “An observer, looking at it, could legitimately say that the molecule ‘behaves’ in a certain way, although in reality [the spider] just implements simple leg residency rules,” says Stojanovic.
Seeman’s group, on the other hand, has managed to get DNA walkers to transport cargo. On their origami, the researchers placed three DNA machines that could be set up to donate or keep different types of cargo, so that a passing walker can take on eight (that is, 23) possible loads and deposit them at the end of the track. “This is important because we have combined a number of elements,” explains Seeman. “The walker, the three independently programmed addition stations and the cargo are all sitting on a DNA origami platform. It is the first assembly line built on the nanometre scale.”
The assembly line produced by Seeman’s group might seem a bigger development than the nibbling spider of Stojanovic’s group. But in fact the latter is just as important because the process is fully autonomous. This is unlike the assembly line, which requires the manual addition of DNA fuel strands to keep the walker moving and accepting cargo. The next step might be to create an autonomous assembly line, just like in biology where independent factories, or “ribosomes”, produce different proteins according to the chemical messages they receive.
“Eventually we want to make something as complex as a cell that can have all these independent little factories running in it, each one cranking out a different product based on their program – and this is what Seeman’s paper is starting to show us how to do,” says Rothermund.
The research is reported in Nature .