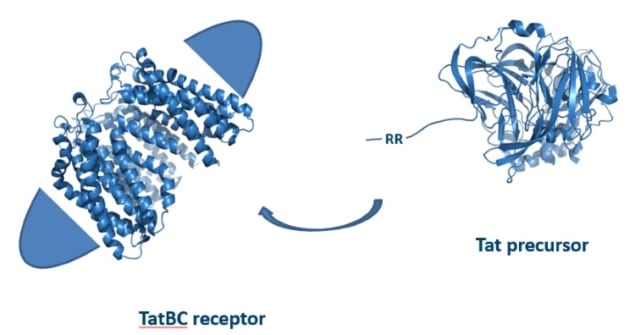
Using a new detergent called glycol-diosgenin (GDN), Marta Wojnowska and her co-workers have managed to investigate a protein complex that secretes proteins. The team, from the University of Oxford, overcame limitations that other researchers had previously faced by solubilizing the protein complex TatBC in the synthetic detergent GDN, rather than the naturally occurring digitonin. Using GDN enabled the use of biophysical analysis methods such as quantitative lipidomics mass spectrometry, isothermal titration calorimetry and surface plasmon resonance (Biochemistry 57 1663).
Foxglove was not good enough
Until recently, the only way to isolate TatBC was by using a detergent called digitonin, which is derived from the flower foxglove. Similar to soap, it disrupts the membrane and solubilizes TatBC. Unfortunately, digitonin also makes biophysical analysis impossible due to “clouding”. Clouding occurs due to detergent instability in solutions over time, which leads to changing background signals during measurements and clogging of instruments.
The new detergent GDN, first synthesized in 2012, was found to be suitable to isolate the TatBC complex. It is a synthetic detergent, but is similar in structure to digitonin. TatBC, isolated using GDN, can bind its substrates with the same specificity as observed in a living cell. The researchers therefore used it to study the thermodynamics and kinetics of interactions between TatBC and the proteins it transports.
TatBC is part of the twin-arginine translocation pathway (Tat) that allows cells to export folded protein. This process presents a challenge, as the cell’s energy is stored in an ion gradient across the cytoplasmic membrane in bacteria, and across the thylakoid membrane in chloroplasts. Leakage of ions during transport depletes the cell’s energy. The proteins TatB and TatC interact to form a complex in the membrane that recognizes the folded substrate protein – ready to be transported -based on its twin-arginine secretion signal. While the structures of TatB and TatC by themselves are known, the TatBC complex could not be studied in isolation in digitonin.
Questions can now be answered
The researchers hoped to answer several questions. How many TatBCs are needed to bind a protein substrate? How tightly and quickly does it bind substrates? Can TatBC work in isolation or is it dependent on other components present in the cell?

Wojnowska, from the research group of Ben Berks, showed that isolated TatBC can bind substrates specifically, but that it does so more slowly than in an intact cell. Two to three TatBCs were present for each substrate protein, either indicating that some TatBCs are inactive or that multiple TatBCs are needed per substrate.
The team also found that the lipid mixture that co-purifies with TatBC is not representative of the overall lipid composition of the membrane TatBC in purified from. As it is known that TatBC is sensitive to different lipid composition, it might be that it is found in areas of the membrane with specific lipid composition or even that it recruits its favourite lipids. However, it might also be that the detergent selectively extracts lipids, which are not ideal for TatBC. This could explain the decreased activity.
A missing piece?
Another explanation for the slow binding could be that a component is missing from the system that is present in whole cells. The researchers now want to use the isolated TatBC complex to investigate whether addition of other cell components, like proteins or lipids, restores quick binding for the isolated protein.