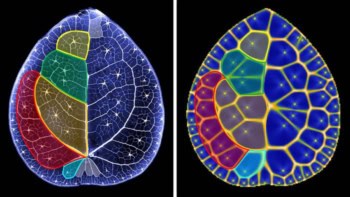
Extremely bright and short-lived pulses of X-rays from a free-electron laser have been used to generate 3D images of virus particles. Unlike existing methods, the technique can be used to identify asymmetries in the structure of biological molecules and could lead to the development of drugs that target molecules whose properties cannot be studied using conventional crystallography.
The X-ray crystallography of molecules involves exposing a lattice of regularly spaced molecules to a beam of X-rays. Interference of the scattered radiation creates a diffraction pattern that is then used to calculate the structure of the constituent molecules.
Although the technique is used today at synchrotrons to carry out a wide range of research in biology, chemistry and physics, it has a weakness: it requires molecules to be assembled into crystals. That leaves out many molecules that are of interest scientifically and practically, including the “membrane proteins” involved in regulating cellular input and output that would need to be targeted by certain drugs.
Ten billion times brighter
This is where X-ray free-electron lasers (XFELs) come into their own. These devices use magnets to direct accelerated electrons along slalom trajectories to generate laser-like pulses of X-rays. The pulses are about 10 billion times brighter than that available at synchrotrons, allowing diffraction patterns to be created from a single molecule – the interference in this case occurring between photons scattered off different points of the molecule. Such intense radiation rips the molecule apart but because the laser pulses are also exceptionally brief – lasting just a few femtoseconds (10–15 s) – most of the diffraction information escapes before the molecule disintegrates.
XFELs, however, struggle to produce 3D images. In a synchrotron, crystals can be placed in the X-ray beam and are then slowly rotated over the course of several hours to generate thousands of diffraction patterns from very slightly different angles. In XFELs, molecules are typically passed through the laser beam inside droplets at a rate of several hundred or thousand a second. The 3D structure can in principle be worked out by passing many copies of the same molecule through the beam one after another. However, the molecules are oriented at random and the challenge is to establish the orientation of each diffraction pattern – a process hampered by noise.
In the latest work, researchers at the SLAC and Lawrence Berkeley laboratories in California and others at the European XFEL near Hamburg, Germany, demonstrate a way round this problem. Rather than determining orientations, their approach exploits what are known as cross-correlation functions. The idea, according to team member Ruslan Kurta of European XFEL, is to correlate intensity values across the diffraction patterns “for all possible combinations of scattering vectors”, in order, he explains, to “work out how those correlations vary as a function of the vector magnitudes and angles between the vectors”. More succinctly, he says, these functions resemble a molecule’s “fingerprint”.
The new research is not the first to use XFELs to make 3D images of virus particles – the genome-carrying part of viruses. Two years ago, a group led by Janos Hajdu of Uppsala University in Sweden employed the Linac Coherent Light Source (LCLS) at SLAC to image 450 :nm-diameter “mimivirus” particles. However, that work relied on the fact that these particles are not only relatively large, but also symmetrical – important because symmetry simplifies 3D reconstruction.
Iterative algorithm
Kurta and colleagues avoid the need for symmetry by using an iterative algorithm to hone the limited information provided by cross-correlation functions. They too put their technique to the test at the LCLS, but used 70 nm-diameter rice dwarf and bacteriophage virus particles. They were able to see nanometre-sized “structural features” of the viruses, including asymmetrical ones. That asymmetry, says team member Adrian Mancuso of European XFEL, might be inherent to the viruses or might instead be an artefact – caused, for example, by the samples getting squashed. But either way, he argues, what matters is that the team has observed asymmetry.
One scientist involved in the previous research, Jean-Michel Claverie of Aix-Marseille University in France, agrees that the latest work represents “sizeable progress” in 3D imaging of single particles. But he cautions that other viruses that might be probed with the new technique – such as the oblong-shaped pandoravirus and pithovirus particles – have quite variable lengths and therefore “might violate” a requirement of the scheme: its need for “reproducible objects”.
Mancuso acknowledges this potential stumbling block, but is confident that it can be overcome. He says that the team now aims to improve the technique’s resolution by using shorter wavelength X-rays and by placing multiple molecules, rather than just a single molecule, in each laser. Using shorter-wavelength X-rays, he adds, should be easier if the group can get beam time on the European XFEL, which opened last month.
The research will be described in an upcoming paper in Physical Review Letters.