A new type of nanotweezer capable of trapping and extracting single entities such as DNA, RNA and mitochondria from a living biological cell could help researchers better understand the fundamentals of cellular processes and how these processes occur in real time. The device, which does not damage cells, unlike many existing such analytical techniques, could also help in the construction of the Human Cell Atlas – the most ambitious genomics project after the sequencing of the human genome.
Many biological cells of the same type – for example brain, muscle or fat cells – look very similar to each other but they have quite different structures and compositions at the single-molecule level. The problem is that conventional approaches to unearth these differences typically require the target cell to be removed from its environment before it can be analyzed. This destroys important information on how the cell contents interconnect, and the process often kills the cell too. Such techniques thus only provide a snapshot of a cell’s transcriptional profile at a given point in time, and cannot provide dynamical information such as molecular changes in the cell as they occur.
Non-destructive technique
A team led by Joshua Edel and Alex Ivanov from Imperial College London has now developed a technique that overcomes this problem. “Our nanotweezers are capable of extracting single molecules from live cells in real time without destroying them,” explains Edel. “We can extract several different parts from different regions of the cell, including mitochondria from the cell body, RNA from different points in the cytoplasm and even DNA from the cell nucleus.”
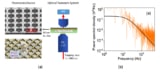
The researchers made their nanotweezers by depositing carbon at the tip of a sharp two-chambered glass capillary to form a pair of electrodes with a 10 to 20 nm insulating gap between them. By applying an alternating current voltage between the electrodes, it is possible to generate a powerful and highly localized electrical field at the tip of the nanotweezers that can be used to trap and extract entities from living cells with single-molecule precision.
“This technique works thanks to an effect known as dielectrophoresis and it could provide us with a deeper understanding of cellular processes – and, for example, find out why cells of the same type can be very different to each other,” say team members Binoy Paulose Nadappuram and Paolo Cadinu. “For example, nerve cells require a lot of energy to fire signals so they contain many mitochondria (which are the “powerhouses” of living cells). By adding or removing mitochondria from individual nerve cells, we could better elucidate their role – particularly in neurodegenerative diseases.”
Testing the nanotweezers
The researchers tested out their nanotweezers by using them to trap and extract small protein- and single DNA molecules from aqueous solutions. They also used them to perform “single-cell biopsies” and extract DNA directly from the nucleus of a human osteosarcoma cell and primary human pulmonary artery endothelial cells. They also succeeded in extracting RNA from the cytoplasm of these cells for subsequent genomic analysis.
And that is not all: the researchers show that the nanotweezers can be used to manipulate single organelles too by trapping and extracting single mitochondria from the hippocampal neurons of mice.
Plasmonic tweezer array traps microparticles
Helping to construct the Human Cell Atlas
“These tweezers are effectively a new tool to interact with single cells and their constituents and have unprecedented spatial resolution, they tell Physics World. “For instance, monitoring cells over time and thus performing multiple biopsies at different points in different locations may help us better understand how cells react to external stimuli and the cellular processes involved in different signaling pathways.”
The team, reporting its work in Nature Nanotechnology 10.1038/s41565-018-0315-8, says that it is now busy integrating the nanotweezers into different scanning probe techniques. “This will allow for spatial and temporal quantification of single gene expression and understanding the role of different organelles in cell function,” say Nadappuram and Cadinu. “The technique could ultimately help in the construction of the Human Cell Atlas, which aims to create a reference map of all human cells.”