Naoki Kunishama and his team at RIKEN’s SPring-8 Center in Japan have found that a new technique to determine protein structures is better for structure-based drug design than conventional X-ray crystallography using synchrotron radiation (SR). The main advantage of this technique, called serial femtosecond crystallography (SFX), is that it can be performed at room temperature without significant noise from radiation damage, while SR relies on cryo-cooling proteins to reduce radiation damage to an acceptable level.
SFX works by exploiting ultrashort pulses of high-energy radiation delivered by an X-ray free-electron laser (XFEL), which is similar in size to a synchrotron but with a linear configuration. To date there are two XFEL facilities, one in California and the one in Japan used by Kunishama and his team, while a third one will soon be operational in Germany.
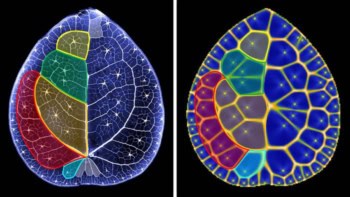
SFX has been optimized for protein crystallography over the last five years. It exploits diffraction from protein crystals, in the same way as synchrotron-based crystallography. Although the radiation used is so intense that it destroys the crystal after a few femtoseconds of exposure, the femtosecond speed of the procedure allows the diffraction data to be captured before radiation damage and the explosion of the crystal can add noise to the data. A full dataset is obtained by working through many crystals each at a fixed random angle, instead of rotating one crystal to take multiple images as in synchrotron techniques. A water- or oil-based stream moves the crystals in line for the radiation pulse.
SFX crystals are more reproducible
In this new study the researchers crystallized thermolysin, a very stable protein that has been used as a model protein for crystallography before, and soaked its ligand into the crystal (Acta Cryst. D73, 702–709). They used SFX to obtain three structures of the complex, and SR for the other two. Comparing the structures to each other, and to previously published structures, Kunishama and colleagues found that SFX yields structures that are more similar to the physiological conditions of both the protein and the water surrounding it. The SFX structures were also much more reproducible than the SR versions.
The big advantages of SFX for generating such highly reproducible structures are that it enables room-temperature operation and generates less radiation damage. In contrast, SR crystallography requires the crystals to be cooled in liquid nitrogen to keep radiation damage within acceptable limits, which in turn needs cryo-protection agents to prevent water crystals from forming inside the protein crystal and destroying it. Kunishama and his team showed that these cryo-protectants affect the conformations of amino acids in the crystal and consequently the observed structure, while cryo-cooling also shrinks the crystal cell dimensions by up to 2.8%.
The devil is in the detail
The authors believe that the more physiological structures obtained by SFX are a better basis for drug design than SR-based structures. Knowing the details of amino-acid conformations is crucial for predicting how drugs can bind to target proteins, especially those concerning the ligand and its binding site, and this study shows that these details might be altered by radiation damage or cryo-cooling in conventional SR crystallography. Kunishama and his team conclude that SFX is the better choice for structure-based drug design, as long as plenty of crystals are available to feed into an SFX crystal stream.