Whole-body positron emission tomography combined with computed tomography (PET/CT) is a cornerstone in the management of lymphoma (cancer in the lymphatic system). PET/CT scans are used to diagnose disease and then to monitor how well patients respond to therapy. However, accurately classifying every single lymph node in a scan as healthy or cancerous is a complex and time-consuming process. Because of this, detailed quantitative treatment monitoring is often not feasible in clinical day-to-day practice.
Researchers at the University of Wisconsin-Madison have recently developed a deep-learning model that can perform this task automatically. This could free up valuable physician time and make quantitative PET/CT treatment monitoring possible for a larger number of patients.
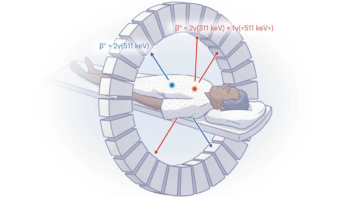
To acquire PET/CT scans, patients are injected with a sugar molecule marked with radioactive fluorine-18 (18F-fluorodeoxyglucose). When the fluorine atom decays, it emits a positron that instantly annihilates with an electron in its immediate vicinity. This annihilation process emits two back-to-back photons, which the scanner detects and uses to infer the location of the radioactive decay.
Because tumours grow faster than most healthy tissue, they must consume more energy. Much of the radioactive tracer will therefore be drawn towards the lymphoma lesions, making them visible in the PET/CT scan. However, other types of tissue, such as certain fatty tissues, can “light up” the scans in a similar manner, which can lead to false positives.
Neural networks: accurate and fast
In their study, published in Radiology: Artificial Intelligence, Amy Weisman and colleagues investigated lesion-identifying deep-learning models built from different configurations of convolutional neural networks (CNNs). They trained, tested and validated these models using PET/CT scans of 90 patients with Hodgkin lymphoma or diffuse large B-cell lymphoma. For this purpose, a single radiologist delineated lesions within each scan and classified each one on a scale from 1–5, depending on how sure they were that a lesion was malignant.
The researchers found that a model consisting of three CNNs performed best, identifying 85% of manually contoured lesions (923 of 1087, the so-called true positive rate). At the same time, it falsely identified four lesions per patient (the false positive rate). The time to evaluate a single scan was cut from 35 minutes using manual delineation to under two minutes for the model.
It is extremely difficult to classify every lymph node in a scan as cancerous or not with 100% certainty. Because of this, if two radiologists delineate lesions for the same patient, they are not likely to agree with each other completely. When a second radiologist evaluated 20 of the scans, their true positive rate was 96%, while they marked on average 3.7 malignant nodes per patient that their colleague had not. In these 20 patients, the deep-learning model had a true positive rate of 90%, at 3.7 false positives per scan – making its predictions almost as good as the variation between two observers.
Expected, and unexpected, challenges
Often, one of the biggest hurdles in creating this type of model is that training it requires a large number of carefully delineated scans. The study authors tested how well their model performed depending upon the number of patients used for training. Interestingly, they found that a model trained on 40 patients performed just as well as one trained on 72.

According to Weisman, obtaining the detailed lesion delineations for training the models proved a more challenging task: “Physicians and radiologists don’t need to carefully segment the tumours, and they don’t need to label a lesion on a scale from 1 to 5 in their daily routine. So asking our physicians to sit down and make decisions like that was really awkward for them,” she explains.
The initial awkwardness was quickly overcome, though, says Weisman. “Because of this, Minnie (one of our physicians) and I got really close during the time she was segmenting for us – and I could just text her and say ‘What was going on with this image/lesion?’. Having a relationship like that was super helpful.”
Future research will focus on incorporating additional, and more diverse, data. “Acquiring more data is always the next step for improving a model and making sure it won’t fail once it’s being used,” says Weisman. At the same time, the group is working on finding the best way for clinicians to use and interact with the model in their daily work.